RESEARCH THEMES

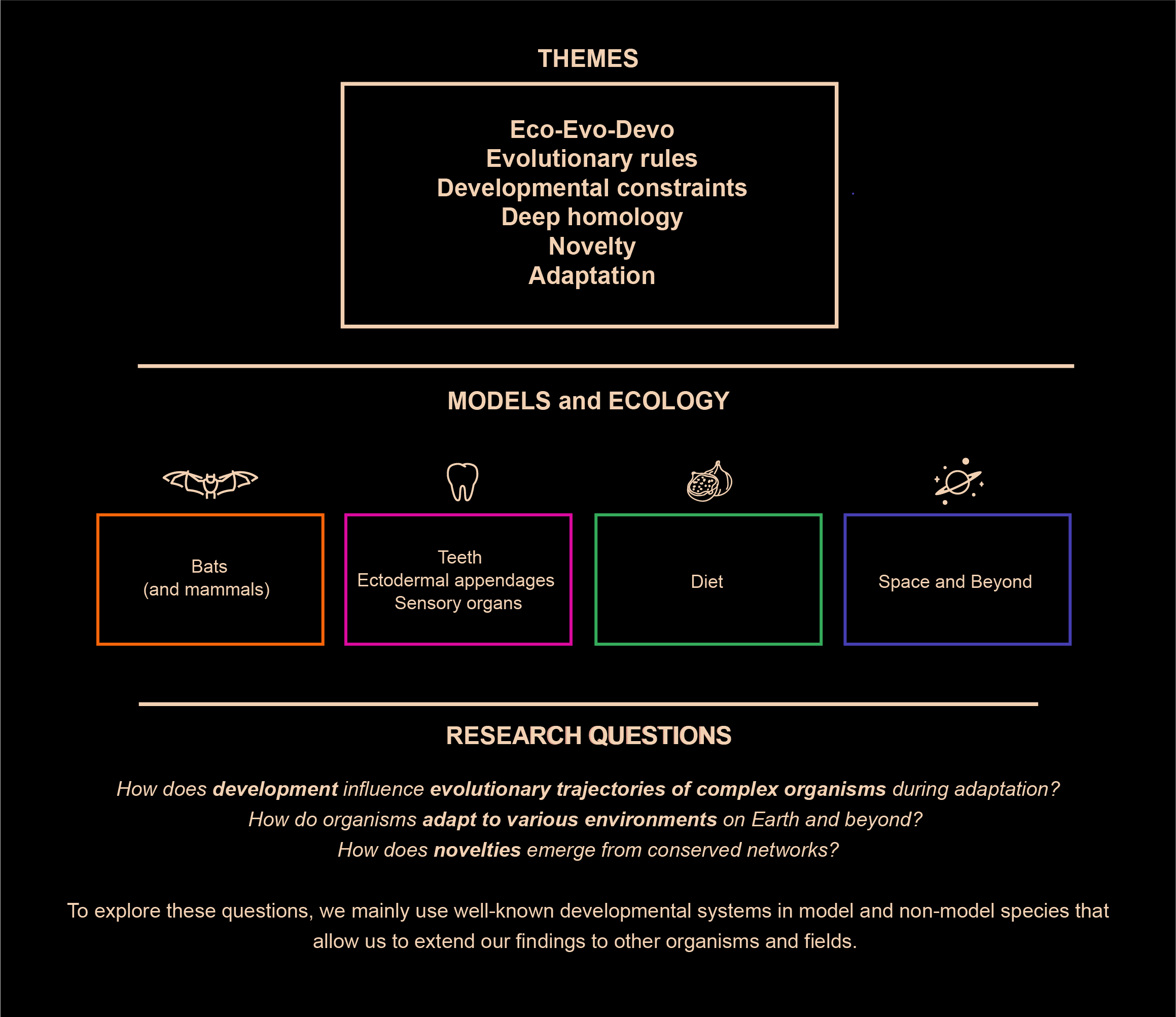
Evolutionary novelties
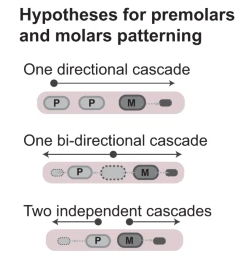
One long-standing question in evolutionary biology is: how do new morphological innovations arise? While ntegrative advances in palaeontology, genomics, development and evolution have unveiled general principles behind evolutionary novelties, we are far from understanding the mechanisms by which novelties emerge and diversify in new ecological contexts. To solve this problem, we use the dramatic morphological and ecological diversity of bat teeth as a model system to study the origin and diversification of a major mammalian innovation, the tooth classes.
Developmental constraints
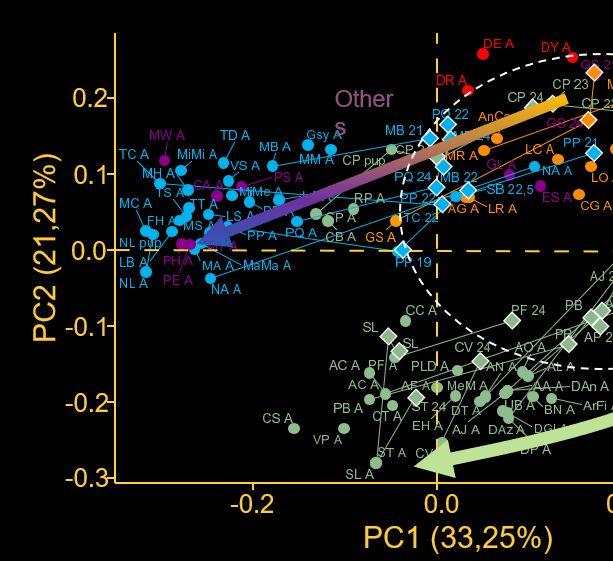
Why some morphologies frequently evolve, while others, though theoretically possible, never arise? As phenotypes are the product of phenotypes through development, this paradox has often been explained by the existence of biases or contraints on developmental processes. While the genomic and ecological drivers of variation in adult morphologies have been a frequent and productive subject of study, how developmental processes facilitate or constrain morphological evolution remains poorly understood. In our lab, we use teeth and ectodermal appendages to decipher how developmental constraints both limit and facilitate the evolution of phenotypes on Earth and beyond.

Sensory adaptations
Bats have evolved specialized sensory systems to forage, hunt and orientate themselves, sometimes in complete darkness. Like many other traits, sensory systems like vision and echolocation have diversified following the colonization of various dietary niches. In the lab, we study the evolution of vision and echolocation using intergative approaches during development. We showed that color vision is mosaic in Noctilionoid bats, UV vision being lost multiple time independently. We also showed that the development of the bat cochlea constraint the development of other sensory systems.
Work related to the origin and diversification of mammalian tooth classes and developmental constraints:
Sadier, A., Anthwal, N., Krause, A.L. et al. Bat teeth illuminate the diversification of mammalian tooth classes. Nat Commun14, 4687 (2023). https://doi.org/10.1038/s41467-023-40158-4
Grossnickle, D. M., Sadier, A., Patterson, E., Cortés-Viruet, N. N., Jiménez-Rivera, S. M., Sears, K. E., & Santana, S. E. (2024). The hierarchical radiation of phyllostomid bats as revealed by adaptive molar morphology. Current Biology. https://doi.org/10.1016/j.cub.2024.02.027
Sadier, A., Twarogowska, M., Steklikova, K., Hayden, L., Lambert, A., Schneider, P., Laudet, V., Hovorakova, M., Calvez, V., and Pantalacci, S. (2019). Modeling Edar expression reveals the hidden dynamics of tooth signaling center patterning. PLoS Biol 17, e3000064 https://doi.org/10.1371/journal.pbio.3000064
Sadier A., Sears, K. and Womack, M. Unravelling the heritage of lost traits, J Exp Part B. https://doi.org/10.1002/jez.b.23030
Work related to bat vision and cochlea evolution:
Anthwal, N., Hall, R. P., de la Rosa Hernandez, F. A., Koger, M., Yohe, L. R., Hedrick, B. P., Davies, K. T. J., Mutumi, G. L., Roseman, C. C., Dumont, E. R., Davalos, L. M., Rossiter, S. J., Sadier*, A. (co-last), & Sears*, K. E. (2023). Cochlea development shapes bat sensory system evolution. The Anatomical Record, 1–12. https://doi.org/10.1002/ar.25353
Sadier*, A., Davies*, K.T., Yohe, L.R., Yun, K., Donat, P., Hedrick, B.P., Dumont, E.R., Davalos, L.M., Rossiter, S.J., and Sears, K.E. (2018). Multifactorial processes underlie parallel opsin loss in neotropical bats. Elife 7. https://elifesciences.org/articles/37412
Work related to bat evolution and ecology:
Sharlene E Santana, Alexa Sadier, Marco A R Mello, (2024). The ecomorphological radiation of phyllostomid bats, Evolutionary Journal of the Linnean Society, https://doi.org/10.1093/evolinnean/kzae032
Sadier A, Urban DJ, Anthwal N, Howenstine AO, Sinha I, Sears KE, Sadier A, Urban DJ, Anthwal N, Howenstine AO, Sinha I, Sears KE. 2020. Making a bat: The developmental basis of bat evolution. Genetics and Molecular Biology http://dx.doi.org/10.1590/1678-4685-gmb-2019-0146
Work related to EDA pathway evolution:
Sadier, A., Lambert, E., Chevret, P., Decimo, D., Semon, M., Tohme, M., Ruggiero, F., Ohlmann, T., Pantalacci, S., and Laudet, V. (2015). Tinkering signaling pathways by gain and loss of protein isoforms: the case of the EDA pathway regulator EDARADD. BMC Evol Biol 15, 129. https://doi.org/10.1186/s12862-015-0395-0
Sadier, A., Viriot, L., Pantalacci, S., and Laudet, V. (2014). The ectodysplasin pathway: from diseases to adaptations. Trends Genet 30, 24–31. https://doi.org/10.1016/j.tig.2013.08.006
